Pancreas 4 Data
Kevin Wang
12 Apr 2024
Source:vignettes/case_study/Pancreas4_Data.Rmd
Pancreas4_Data.RmdIntroduction
This is a human pancreas single-cell data comprising of 4 experiments.
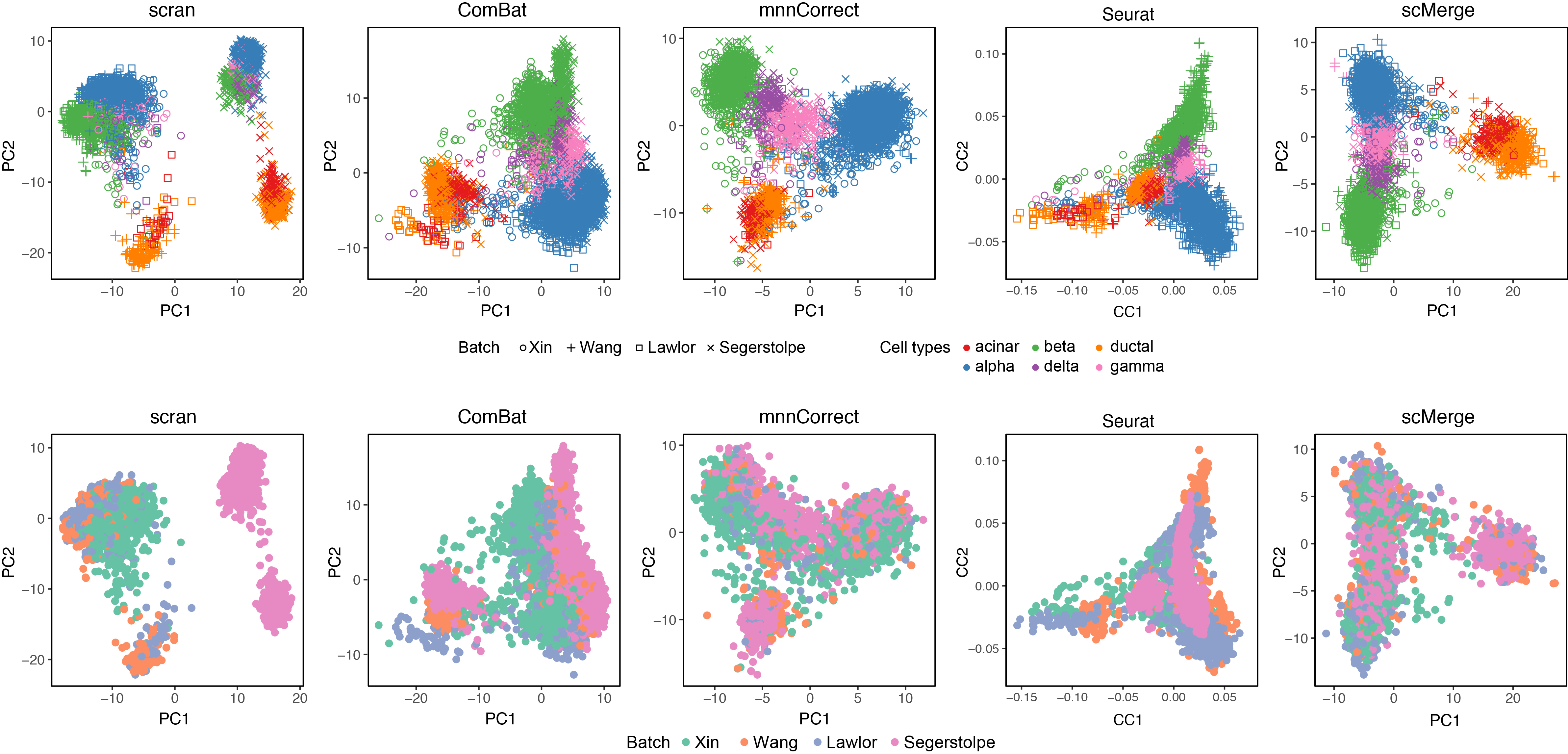
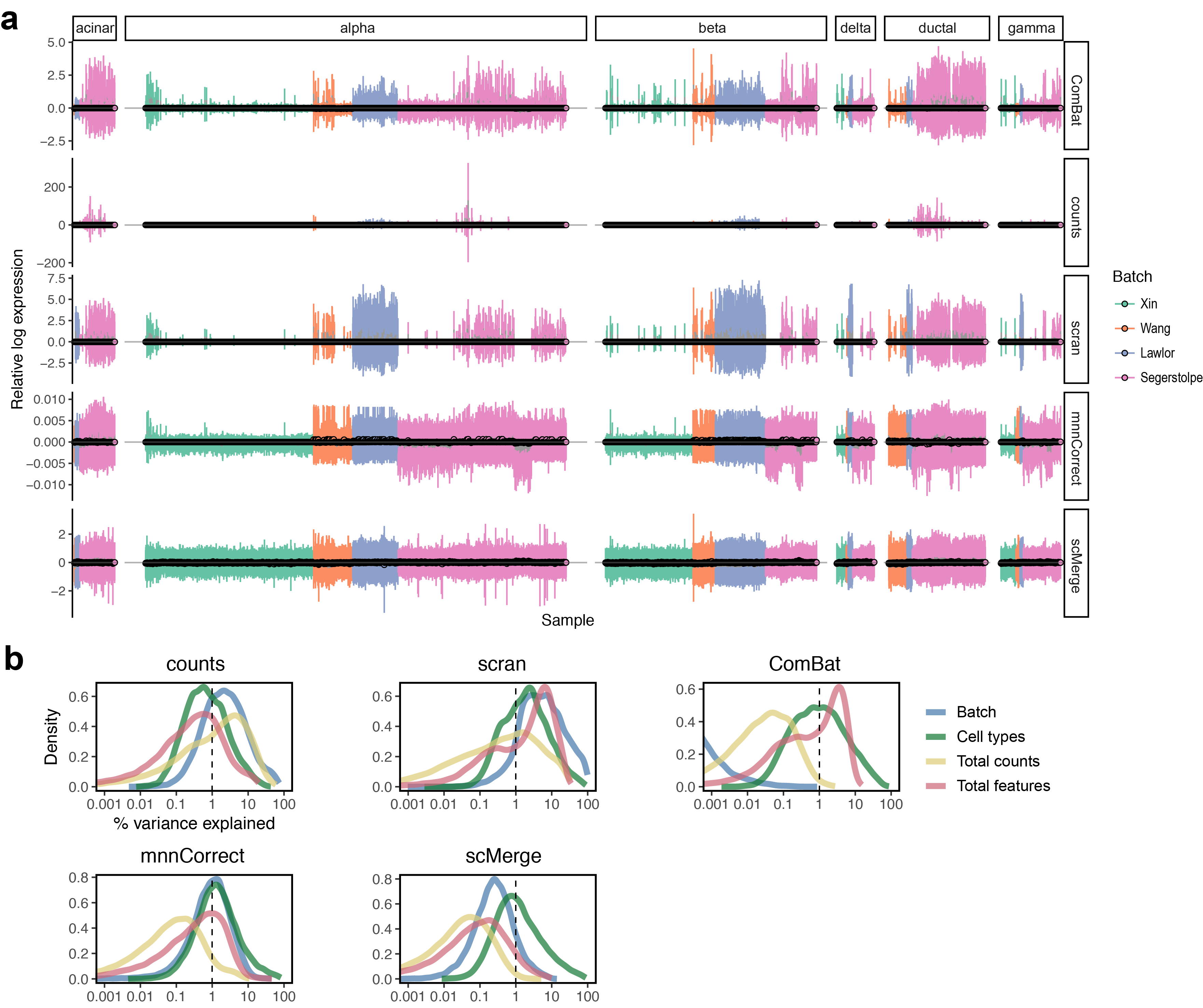
Integration challenge
- Prior to integration, there is a strong separation effect by experiments.
- In addition, there are 3 different protocols, the sequencing depths poses an extra challenge.
- With 4,566 cells, this data poses a moderate dimensionality challenge to data integration.
Data description
- Data source:
| Type of merge | Name | ID | Author | DOI or URL | Protocol | Organism | Tissue | # of cell types | # of cells | # of batches | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Across platforms with significant depth difference | Pancreas4 | GSE81608 | Xin | 10.1016/j.cmet.2016.08.018 | SMARTer/C1 | Human | Pancreas | 6 | 4566 | NA | |
| GSE83139 | Wang | 10.2337/db16-0405 | Smart-Seq | ||||||||
| GSE86469 | Lawlor | 10.1101/gr.212720.116 | SMARTer/C1 | ||||||||
| E-MTAB-5061 | Segerstolpe | 10.1016/j.cmet.2016.08.020 | Smart-Seq2 |
- Relation to the
scMergearticle: Supplementary Figure 4 and 5.
Integrated scMerge data
Data availability: Pancreas4 (in RData format)
-
scMergeparameters for integration:- Unsupervised scMerge
- kmeans K = (4,6,6,6)
- Negative controls are human scSEG